The mechanism and function of RNAi: nuclear RNAi (Nrde) inheritance of RNAi Germ granule
The struture and regulation of nucleoli: antisense ribosomal siRNA (risiRNA) nucleolar reshaping
The biogenesis of piRNAs: piRNA biogenesis
Aging and epigenetics: aging and epigenetics
Technology: CRISPR
Our lab has been focusing on the mechanism and function of small regulatory RNAs in C. elegans and has discovered the nuclear RNAi defective (Nrde) pathway, a new class of antisense ribosomal siRNAs (risiRNA) and nucleolar RNAi. We also investigated the protein machineries for piRNA transcription and processing and found the upstream sequene transcription complex (USTC), the piRNA biogenesis and chromosome segregation complex (PICS) and a new H3K27me3 reader UAD-2 which is essential for piRNA transcription. We examined the artichecture of germ granules and ideintifed the E- and D- granules which are important for small RNA production. Meanwhile, we investigated the mechanism of transgenerational inheritance of RNAi and found that both the nuclear and cytoplasmic RNAi machineries are involved in the inheritance of RNAi. In addition, we are interested in the structure regulation of nucleolus and found that rRNA interemediate could reshape the nucleolus. In addition, we developed methods using CRISPR-Cas9 technology to induce deletion of large genome fragments, chromosome translocation and inversion.
The mechanistic underpinnings of RNAi are broadly conserved across eukaryotes. Initial successes utilizing small RNAs to target oncogenic and viral mRNAs have generated excitement that small RNAs may eventually be utilized to treat human diseases. Prior to the rational use of small RNAs in therapeutics, it is essential to understand their bi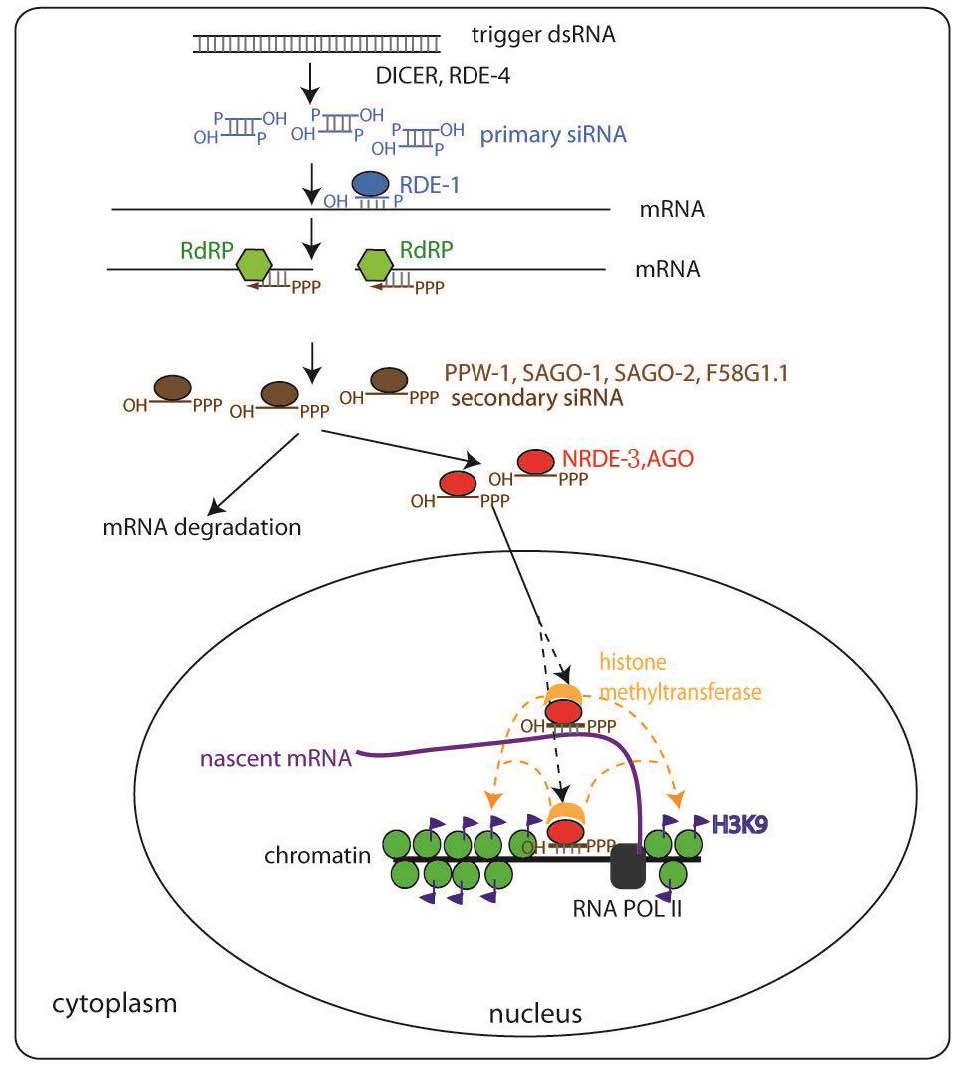 ogenesis, specificity, transportation, and endogenous roles. (Science 2008; Nature 2010; PLoS Genetics 2011; Nature Genetics 2012; Genetics 2014; Current Biology 2015; SCIENCE CHINA Life Sciences 2025a)
ogenesis, specificity, transportation, and endogenous roles. (Science 2008; Nature 2010; PLoS Genetics 2011; Nature Genetics 2012; Genetics 2014; Current Biology 2015; SCIENCE CHINA Life Sciences 2025a)
(1) The identification of nuclear RNAi in C. elegans.
We are very interested in how small RNAs are transported and regulated, andhow they function in the nucleus in metazoan. To address these questions, we conducted a genetic screen to identify factors required for nuclear RNAi in the model organism C. elegans and have identified three new genes termed nuclear RNAi defective (NRDE)-1/2/3. We have also identified a bifurcation of the nuclear and cytoplasmic RNAi pathway, a novel small RNA transport pathway, and a novel nucleargene silencing mechanism.
NRDE-3 is an Argonaute. Argonaute proteins are known to bind small RNAs recognizing complementary cellular RNAs via Watson-Crick base pairing and inhibit gene expression by a variety of mechanisms. Our work revealed that these Argonaute proteins can also escort small RNAs to their distinct subcellular compartments to silence target genes. In the absence of small RNAs, NRDE-3 resides in the cytoplasm. NRDE-3 transports small RNAs from the cytoplasm to the nucleus and associates with nascent transcripts generated by RNA polymerase II. This is the first demonstration that a specific Argonaute protein has transportation activity in addition to silencing.
(2) Nuclear RNAi mediates pre-mature transcription termination.
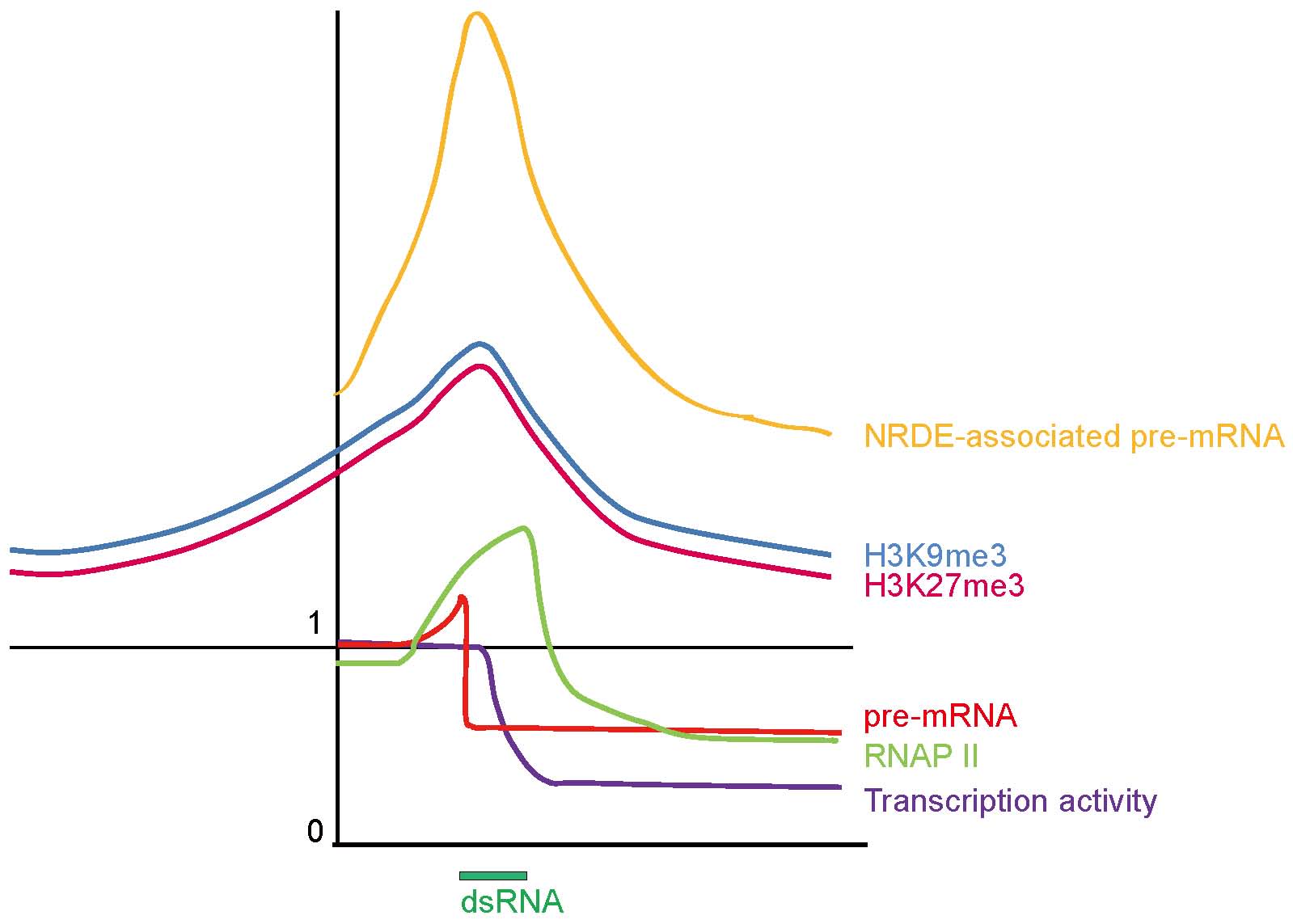 Our work also established a novel nuclear gene silencing pathway in which small RNAs act in conjunction with these newly identified NRDE genes to terminate RNA polymerase II-mediated transcription by pausing the polymerase during its elongation phase. By combining a variety of genetic and biochemical assays, we have found a small RNA and NRDE-dependent silencing of pre-mRNAs 3’ to sites of RNAi, accumulation of RNA Polymerase II at genomic region targeted by RNAi, and decreases in RNA polymerase II occupancy and transcription activity 3’ to sites of RNAi. These experiments demonstrated that metazoan small RNAs elicit a co-transcriptional gene-silencing program, and can act as trans-acting terminators of RNA polymerase II.
Our work also established a novel nuclear gene silencing pathway in which small RNAs act in conjunction with these newly identified NRDE genes to terminate RNA polymerase II-mediated transcription by pausing the polymerase during its elongation phase. By combining a variety of genetic and biochemical assays, we have found a small RNA and NRDE-dependent silencing of pre-mRNAs 3’ to sites of RNAi, accumulation of RNA Polymerase II at genomic region targeted by RNAi, and decreases in RNA polymerase II occupancy and transcription activity 3’ to sites of RNAi. These experiments demonstrated that metazoan small RNAs elicit a co-transcriptional gene-silencing program, and can act as trans-acting terminators of RNA polymerase II.
Taken together, this research indicates that metazoans use a different mechanism other than A. thaliana and S. pombe to silence gene expression in the nucleus. Further understanding how small RNAs function in the nucleus via NRDEs may permit more stable and specific inhibition of gene expression and facilitate advancement of both basic research and therapeutics.
(3) Nuclear RNAi-induced epigenetic modifications.
Ribonucleoprotein co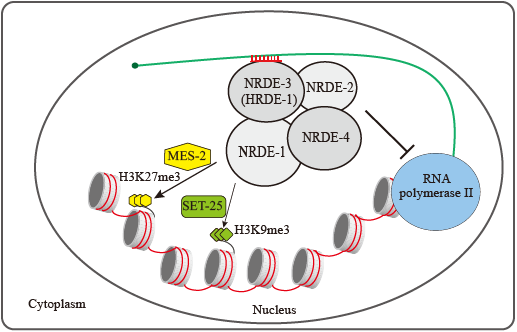 mplexes consisting of Argonaute-like proteins and small regulatory RNAs function in a wide range of biological processes. Many small regulatory RNAs are thought to function, at least in part, within the nucleus to regulate transcription and modify chromatin. We showed that dsRNA triggers H3K27 trimethylation in a sequence-dependent manner, which can be maintained and inherited to progenies for multiple generations. This modification requires the Nrde pathway in Caenorhabditis elegans. Endogenous small RNAs, including, but not limited to, NRDE-3 and HRDE-1-associated endo-siRNAs, induce H3K27me3 in an NRDE-dependent manner. Small RNA-mediated H3K9me3 and H3K27me3 have distinct genetic requirements and different roles in RNAi. The H3K27 methyltransferase, mes-2, is likely to be involved in small RNA-induced H3K27me3.
mplexes consisting of Argonaute-like proteins and small regulatory RNAs function in a wide range of biological processes. Many small regulatory RNAs are thought to function, at least in part, within the nucleus to regulate transcription and modify chromatin. We showed that dsRNA triggers H3K27 trimethylation in a sequence-dependent manner, which can be maintained and inherited to progenies for multiple generations. This modification requires the Nrde pathway in Caenorhabditis elegans. Endogenous small RNAs, including, but not limited to, NRDE-3 and HRDE-1-associated endo-siRNAs, induce H3K27me3 in an NRDE-dependent manner. Small RNA-mediated H3K9me3 and H3K27me3 have distinct genetic requirements and different roles in RNAi. The H3K27 methyltransferase, mes-2, is likely to be involved in small RNA-induced H3K27me3.
(4) Peri-centrosomal localization of small interfering RNAs.
We generated a GFP-NRDE-3 knock-in transgene through CRISPR/Cas9 technology and observed that NRDE-3 formed peri-centrosomal foci neighboring the tubulin protein TBB-2, other centriole proteins and pericentriolar material (PCM) components in C. elegans embryos. The peri-centrosomal accumulation of NRDE-3 depends on RNA-dependent RNA polymerase (RdRP)-synthesized 22G siRNAs and the PAZ domain of NRDE-3, which is essential for siRNA binding. Mutation of eri-1, ergo-1, or drh-3 significantly increased the percentage of pericentrosome-enriched NRDE-3. At the metaphase of the cell cycle, NRDE-3 was enriched in both the peri-centrosomal region and the spindle. Moreover, the integrity of centriole proteins and pericentriolar material (PCM) components is also required for the peri-centrosomal accumulation of NRDE-3.
Antisense ribosomal siRNA (risiRNA) and nucleolar RNAi
For a long time, small fragments of ribosomal RNA have been widely considered by the research community as non-specific degradation products and neglected as garbage sequence. Little is known about their biological processes, functions and regu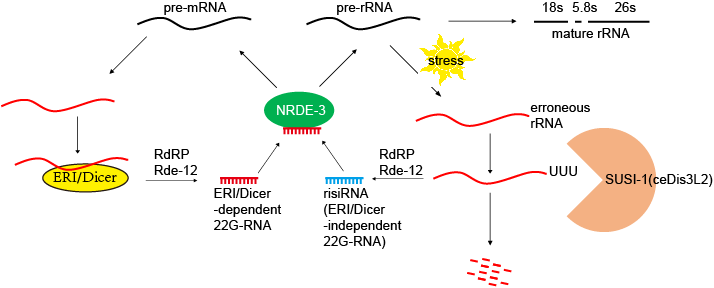 latory mechanisms. Our team found that a new class of antisense ribosomal siRNAs (risiRNAs) downregulate pre-rRNA through a nucleolar RNAi pathway. Interestingly, the nucleolar RNAi pathway also involves the NRDE factors which are essential for nuclear RNAi. (Nature Structural & Molecular Biology 2017; PNAS 2018; Nucleic Acids Research 2021)
latory mechanisms. Our team found that a new class of antisense ribosomal siRNAs (risiRNAs) downregulate pre-rRNA through a nucleolar RNAi pathway. Interestingly, the nucleolar RNAi pathway also involves the NRDE factors which are essential for nuclear RNAi. (Nature Structural & Molecular Biology 2017; PNAS 2018; Nucleic Acids Research 2021)
The risiRNAs are sensitive to environmental stimuli and gene mutations.we conducted both forward and reverse
genetic screens to search for more suppressor of siRNA (susi) mutants and isolated a number of genes that are broadly conserved from yeast
to humans and are involved in pre-rRNA modification and processing. Mutation of
these genes led to a deficiency in modification of rRNAs and elicited
accumulation of risiRNAs, which further triggered the cytoplasmsm to
nucleus and cytoplasmsm to nucleolus translocations of the Argonaute
protein NRDE-3. When the genes SUSI-1(ceDis3L2) and exosomes, which are involved in rRNA degradation, are mutated, risiRNAs are also dramatically increased. Interesting, this SUSI-1(ceDis3L2) mutation is also identified in a human disease called Perlman's syndrome. Therefore, we concluded
that erroneous rRNAs can trigger risiRNA generation and subse-
quently, turn on the nuclear RNAi-mediated gene silencing pathway
to inhibit pre-rRNA expression, which may provide a quality control
mechanism to maintain homeostasis of rRNAs.
In the presence of risiRNAs, both NRDE-2 and NRDE-3 accumulated in the nucleolus and colocalized with RNA polymerase I and associate with pre-rRNA. risiRNAs inhibited the transcription elongation of RNA polymerase I by decreasing RNAP I occupancy downstream of the RNAi-targeted site.
The mechanism of transgenerational inheritance of RNAi.
RNAi-elicited gene silencing is heritable and can persist for multiple generation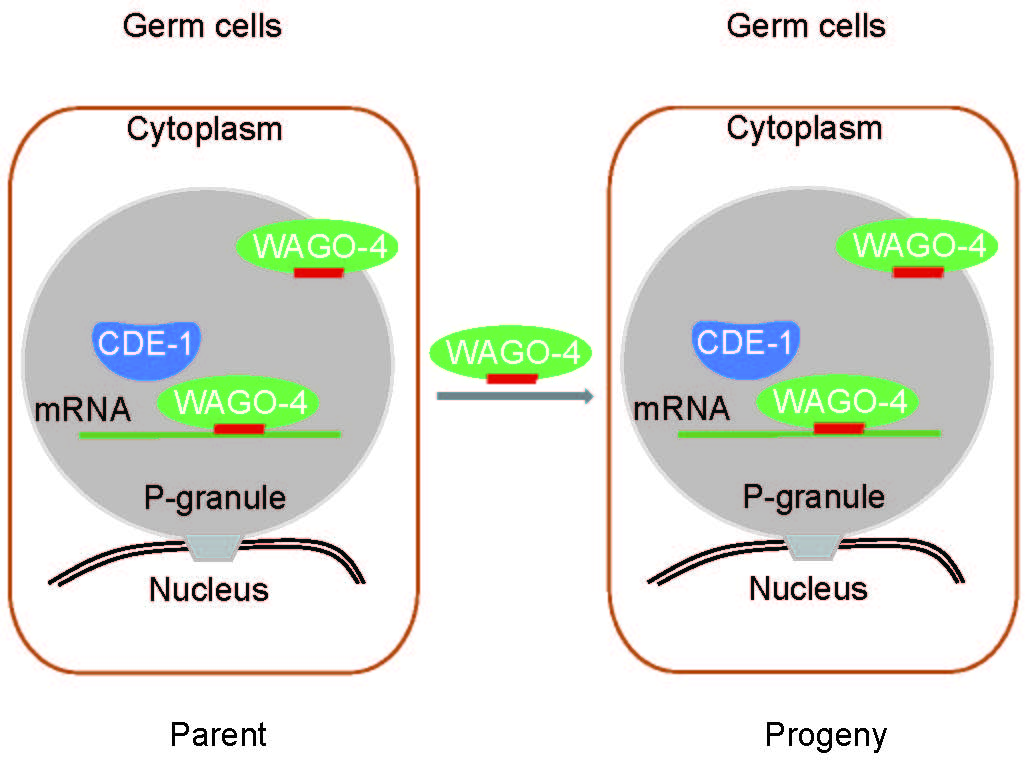 s after its initial induction in C. elegans. However, the mechanism by which parental-acquired trait-specific information from RNAi is inherited by the progenies is not fully understood. We identified a cytoplasmic Argonaute protein, WAGO-4, necessary for the inheritance of RNAi. WAGO-4 exhibits asymmetrical translocation to the germline during early embryogenesis, accumulates at the perinuclear foci in the germline, and is required for the inheritance of exogenous RNAi targeting both germline- and soma-expressed genes. WAGO-4 binds to 22G-RNAs and their mRNA targets. Interestingly, WAGO-4-associated endogenous 22G-RNAs target the same cohort of germline genes as CSR-1 and contain untemplated addition of uracil at the 3' ends. The poly(U) polymerase CDE-1 is required for the untemplated uridylation of 22G-RNAs and inheritance of RNAi. Therefore, we conclude that, in addition to the nuclear RNAi pathway, the cytoplasmic RNAi machinery also promotes RNAi inheritance. (Nature Genetics 2012; Cell Reports 2018)
s after its initial induction in C. elegans. However, the mechanism by which parental-acquired trait-specific information from RNAi is inherited by the progenies is not fully understood. We identified a cytoplasmic Argonaute protein, WAGO-4, necessary for the inheritance of RNAi. WAGO-4 exhibits asymmetrical translocation to the germline during early embryogenesis, accumulates at the perinuclear foci in the germline, and is required for the inheritance of exogenous RNAi targeting both germline- and soma-expressed genes. WAGO-4 binds to 22G-RNAs and their mRNA targets. Interestingly, WAGO-4-associated endogenous 22G-RNAs target the same cohort of germline genes as CSR-1 and contain untemplated addition of uracil at the 3' ends. The poly(U) polymerase CDE-1 is required for the untemplated uridylation of 22G-RNAs and inheritance of RNAi. Therefore, we conclude that, in addition to the nuclear RNAi pathway, the cytoplasmic RNAi machinery also promotes RNAi inheritance. (Nature Genetics 2012; Cell Reports 2018)
The mechanism of piRNA biogenesis
Piwi-interacting RNAs (piRNAs) engage Piwi proteins to suppress transposons and non-self nucleic acids, maintain genome integrity, and are essential for fertility in a variety of organisms. In C. elegans most piRNA precursors are transcribed from two genomic clusters that contain thousands of individual piRNA transcription units. While a few genes have been shown to be required for piRNA biogenesis the mechanism of piRNA transcription remains elusive. We used forward genetic screening together with functional proteomics approaches to identify an upstream sequence transcription complex (USTC), a piRNA and chromosomal segregation (PICS) complex and a new H3K27me3 reader UAD-2 that are essential for piRNA biogenesis. (Genes & Development 2019; Cell Reports 2019; PNAS 2021; Nature Communications 2021, 2023; Journal of Genetics and Genomics 2022; Nature Structural & Molecular Biology 2025; Nature Communications 2025b)
(1) piRNA transription
The USTC complex contains PRDE-1, SNPC-4, TOFU-4 and TOFU-5. The USTC complex form a unique piRNA foci in germline nuclei and coat the piRNA cluster genomic loci. USTC factors associate with the Ruby motif just upstream of type I piRNA genes.
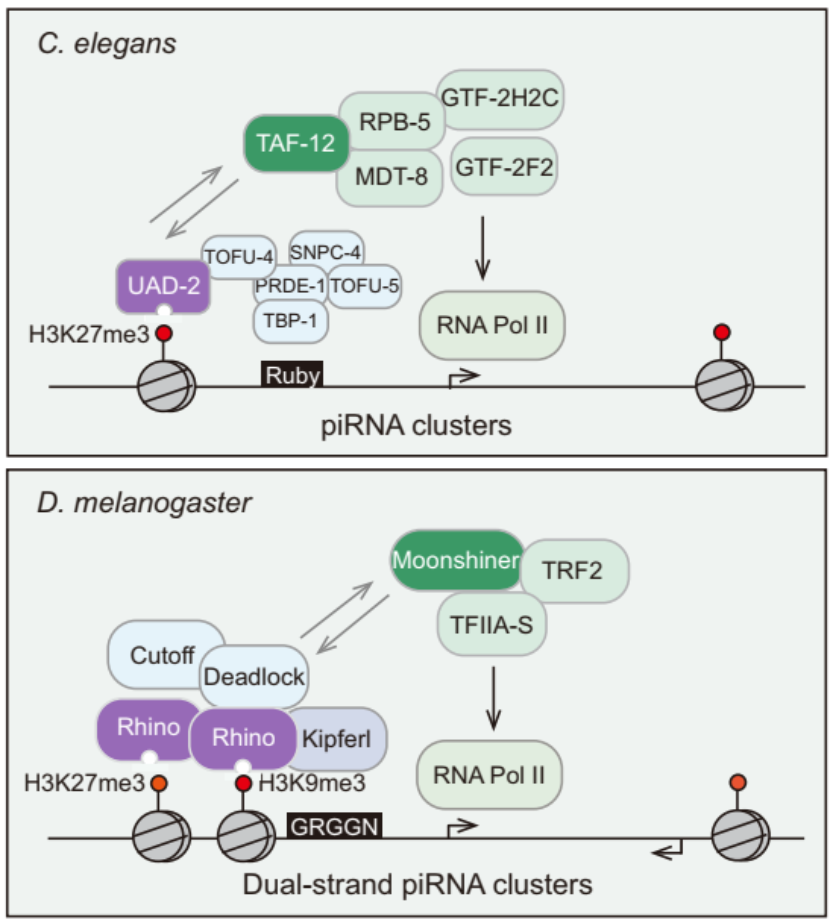
We conducted a forward genetic screening and identified UAD-2 that is required for piRNA focus formation. In the absence of histone 3 lysine 27 methylation or proper chromatin-remodeling status, UAD-2 is depleted from the piRNA focus. UAD-2 recruits the upstream sequence tran scription complex (USTC), which binds the Ruby motif to piRNA promoters and promotes piRNA generation. Vice versa, the USTC complex is required for UAD-2 to associate with the piRNA focus. Thus, transcription of heterochromatic small RNA source loci relies on coordinated recruitment of both the readers of histone marks and the core transcriptional machinery to DNA. Interestingly, the UAD-2 could recruit the core transription factor TAF-12 to initiate piRNA transcription.
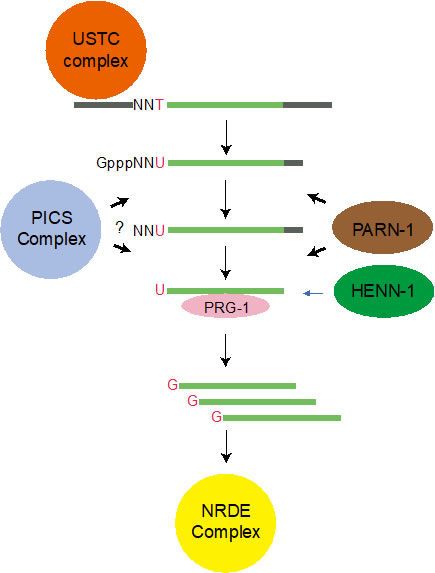
(2) piRNA maturation
We use functional proteomics in C. elegans and identify a piRNA biogenesis and chromosome segregation (PICS) complex. The PICS complex contains TOFU-6, PID-1, PICS-1, TOST-1, and ERH-2, which exhibit dynamic localiza tion among different subcellular ompartments. In the germlines, the PICS complex contains TOFU-6/PICS-1/ERH-2/PID-1, is largely concentrated at the perinuclear granule zone, and engages in piRNA pro cessing. During embryogenesis, the TOFU-6/PICS-1/ERH-2/TOST-1 complex accumulates in the nucleus and plays essential roles in chromosome segregation. The functions of these factors in medi ating chromosome segregation are independent of piRNA production. We speculate that differential compositions of PICS factors may help cells coordinate distinct cellular processes.
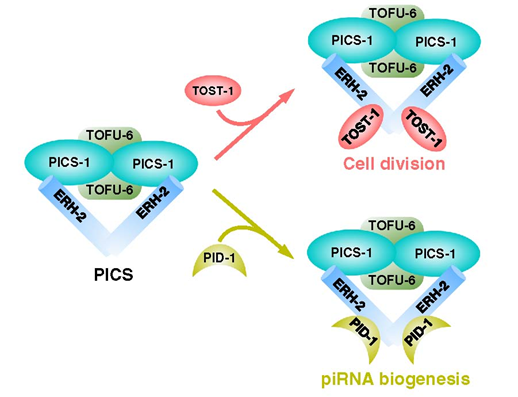
We map the interaction network between PICS subunits, then uncover the mechanisms underlying the interactions between PICS subunits by solving several complex structures, including those of TOFU-6/PICS-1, ERH-2/PICS-1, and ERH-2/TOST-1. Our biochemical experiment also demonstrates that PICS exists as an octamer consisting of two copies of each subunit. Combining structural analyses with mutagenesis experiments, we identify interfacial residues of PICS subunits that are critical for maintaining intact PICS complex in vitro. Furthermore, using genetics, cell biology and imaging experiments, we find that those mutants impairing the in vitro interaction network within PICS, also lead to dysfunction of PICS in vivo, including mislocalization of PICS, and reduced levels of piRNAs or aberrant chromosome segregation and cell division.
Germ granules are biomolecular condensates present in most animal germ cells. One function of germ granules is to help maintain germ cell totipotency by organizing mRNA regulatory machinery, including small RNA-based gene regulatory pathways. The C. elegans germ granule is compartmentalized into multiple subcompartments whose biological functions are largely unknown. Here, we identify an uncharted subcompartment of the C. elegans germ granule, which we term the E granule. The E granule is nonrandomly positioned within the germ granule. We identify five proteins that localize to the E granule, including the RNA-dependent RNA polymerase (RdRP) EGO-1, the Dicer-related helicase DRH-3, the Tudor domain-containing protein EKL-1, and two intrinsically disordered proteins, EGC-1 and ELLI-1. Localization of EGO-1 to the E granule enables synthesis of a specialized class of 22G RNAs, which derive exclusively from 5’ regions of a subset of germline-expressed mRNAs. Defects in E granule assembly elicit disordered production of endogenous siRNAs, which disturbs fertility and the RNAi response. Our results define a distinct subcompartment of the C. elegans germ granule and suggest that one function of germ granule compartmentalization is to facilitate the localized production of specialized classes of small regulatory RNAs. (Nature Communications 2024a; Science China Life Sciences 2025b)
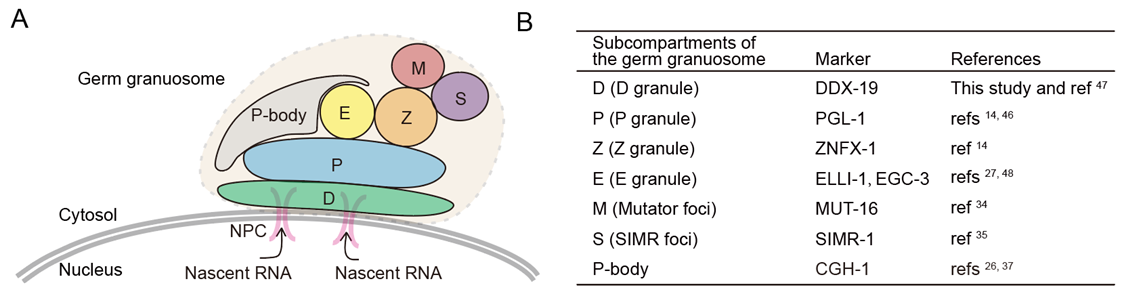
we systematically labeled perinuclear proteins with fluorescent tags via CRISPR/Cas9 technology. Using this nematode strain library, we identified a series of proteins localized in Z or E granules and extended the characterization of the D granule. Finally, we found that the LOTUS domain protein MIP-1/EGGD-1 regulated the multiphase organization of the germ granule. Overall, our work identified the germ granule architecture and redefined the compartmental localization of perinuclear proteins. (Developmental Cell 2024)
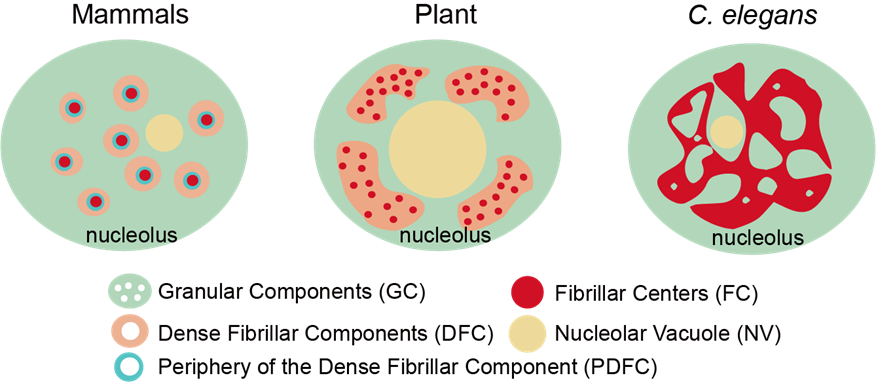
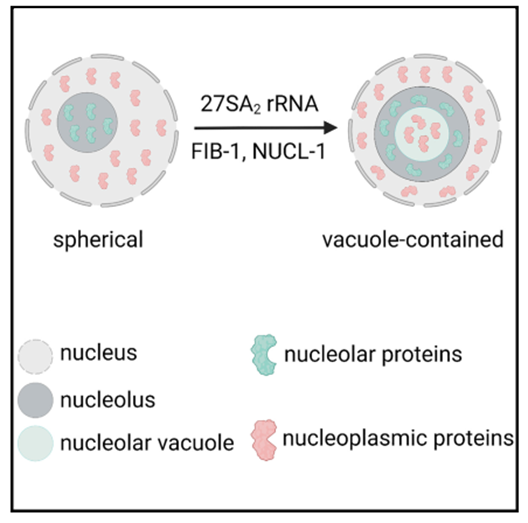
The nucleolus is the most prominent membraneless organelle within the nucleus. How the nucleolar structure is regulated is poorly understood. Here, we identified two types of nucleoli in C. elegans. Type I nucleoli are spherical and do not have visible nucleolar vacuoles (NoVs), and rRNA transcription and processing factors are evenly distributed throughout the nucleolus. Type II nucleoli contain vacuoles, and rRNA transcription and processing factors exclusively accumulate in the periphery rim. The NoV contains nucleoplasmic proteins and is capable of exchanging contents with the nucleoplasm. The high-order structure of the nucleolus is dynamically regulated in C. elegans. Faithful rRNA processing is important to prohibit NoVs. The depletion of 27SA 2 rRNA processing factors resulted in NoV formation. The inhibition of RNA polymerase I (RNAPI) transcription and depletion of two conserved nucleolar factors, nucleolin and fibrillarin, prohibits the formation of NoVs. This finding provides a mechanism to coordinate structure maintenance and gene expression. (PLoS Genetics 2023; Cell Reports 2023; Mol & Cell Proteomics 2025)
Environmental stimuli not only alter gene expression profiles but also induce structural changes in cells. How distinct nuclear bodies respond to cellular stress is poorly understood. Here, we identify a subnuclear organelle named the nucleolar stress body (NoSB), the formation of which is induced by the inhibition of rRNA transcription or inactivation of rRNA processing and maturation in C. elegans. NoSB does not colocalize with other previously described subnuclear organelles. We conduct forward genetic screening and identify a bZIP transcription factor, named nucleolar stress response-1 (NOSR-1), that is required for NoSB formation. The inhibition of rRNA transcription or inactivation of rRNA processing and maturation increases nosr-1 expression. By using transcriptome analysis of wild-type animals subjected to different nucleolar stress conditions and nosr-1 mutants, we identify that the SR-like protein NUMR-1 (nuclear localized metal responsive) is the target of NOSR-1. Interestingly, NUMR-1 is a component of NoSB and itself per se is required for the formation of NoSB. We conclude that the NOSR-1/NUMR-1 axis likely responds to nucleolar stress and mediates downstream stress-responsive transcription programs and subnuclear morphology alterations in C. elegans.(Nature Communications 2024b)
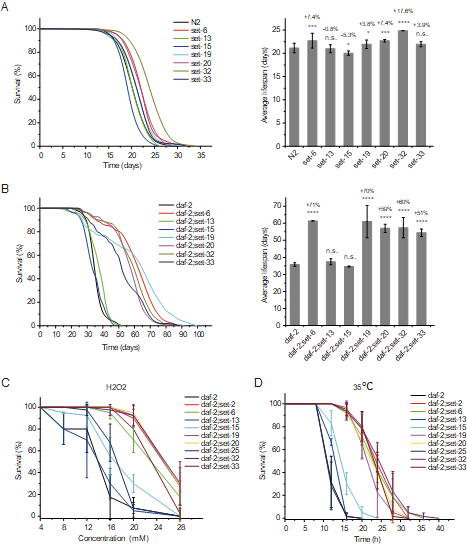
Histone methylation plays crucial roles in the development, gene regulation, and maintenance of stem cell pluripotency in mammals. Recent work shows that histone methylation is associated with aging, yet the underlying mechanism remains unclear. In this work, we identified a class of putative histone 3 lysine 9 mono/dimethyltransferase genes (met- 2, set- 6, set- 19, set- 20, set- 21, set- 32, and set- 33), mutations in which induce synergistic lifespan extension in the long-lived DAF-2 (insulin growth factor 1 [IGF-1] receptor) mutant in Caenorhabditis elegans. These putative histone methyltransferase plus daf- 2 double mutants not only exhibited an average lifespan nearly three times that of wild-type animals and a maximal lifespan of approximately 100 days, but also significantly increased resistance to oxidative and heat stress. Synergistic lifespan extension depends on the transcription factor DAF-16 (FOXO). mRNA-seq experiments revealed that the mRNA levels of DAF-16 Class I genes, which are activated by DAF-16, were further elevated in the daf- 2;set double mutants. Among these genes, tts- 1, F35E8.7, ins- 35, nhr- 62, sod- 3, asm- 2, and Y39G8B.7 are required for the lifespan extension of the daf- 2;set- 21 double mutant. In addition, treating daf- 2 animals with the H3K9me1/2 methyltransferase G9a inhibitor also extends lifespan and increases stress resistance. Therefore, investigation of DAF-2 and H3K9me1/2 deficiency-mediated synergistic longevity will contribute to a better understanding of the molecular mechanisms of aging and therapeutic applications.(eLife 2022; Nature Communications 2023)
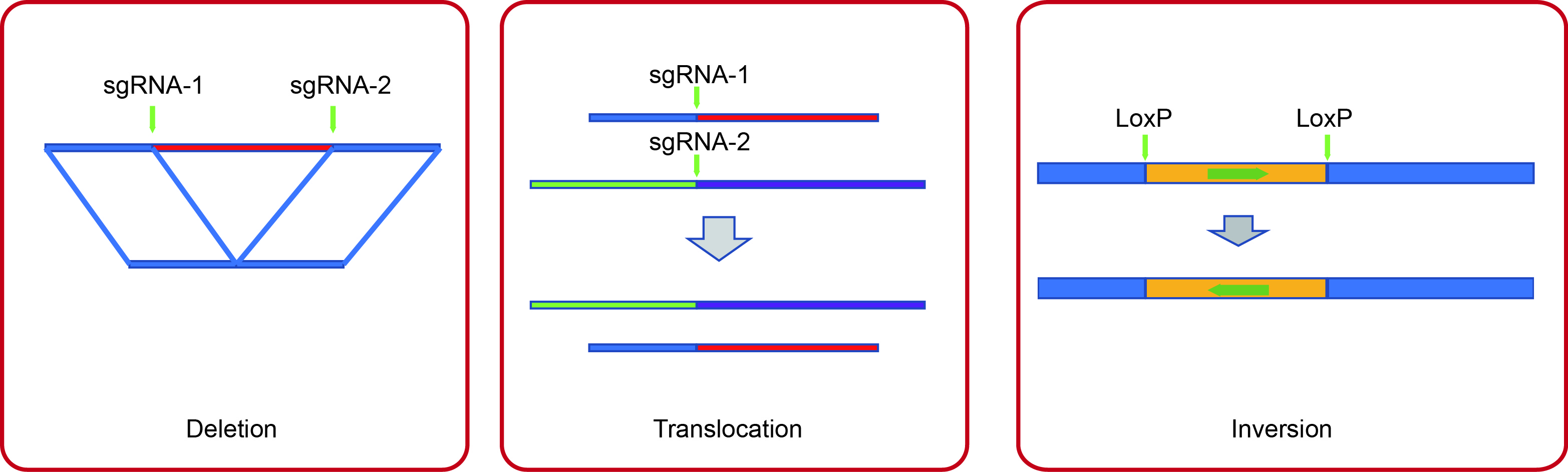
Developing CRISPR/Cas9 technologies for genome manipunation in C. elegans.
The CRISPR/Cas9 technology has been successfully applied for gene editing and chromosome engineering. In the past few years, We have developed the CRISPR/Cas9 techlogies to (a) delete large genomic fragments via dual sgRNAs; (b) induce chromsomal translocations using sgRNAs targeting different chromosomals; (c) induce chrosomal inversions covering thepairing centers by the combinational usage of CRISPR/Cas9 and Cre/LoxP means; and (d) generate a protocol to knock out essential genes in C. elegans. (Scientific Reports 2014; Genetics 2015; G3 2018)